Biolog Lab Services
For over 25 years, our lab has offered comprehensive microbial identification and confirmatory testing with unparalleled customer service. Whether you are concerned about contamination or enumeration, we will work with you to reduce costs and guarantee brand safety.
Biolog Lab Services new address:
225 Corporate Blvd., Suite E, Newark, DE 19702
Trusted Results
ISO 17025: 2017 accredited and validated testing services. Registered with the FDA, GDUFA, and cGMP compliant.
Best in Class
Fastest standard turnaround service ⌄
- Unparalleled turnaround times: 3 days standard
- Same-day results available Monday through Friday
- Fast and concise reports
See individual ID Services for specific turnaround times.
Personal Service
Technical experts are always on hand to answer questions
Quick answers means less time waiting
Flexible Pricing
Bulk, and standing order pricing available for our most popular services. Email us to find out more.
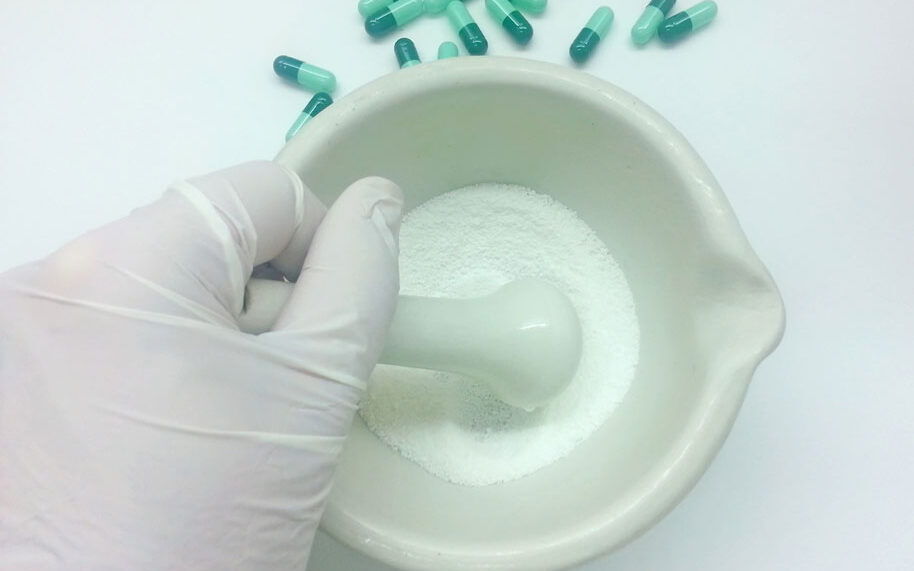

CSPs
Compounding sterile preparations must be in compliance with USP <797> standards. We help you ensure you maintain compliance.
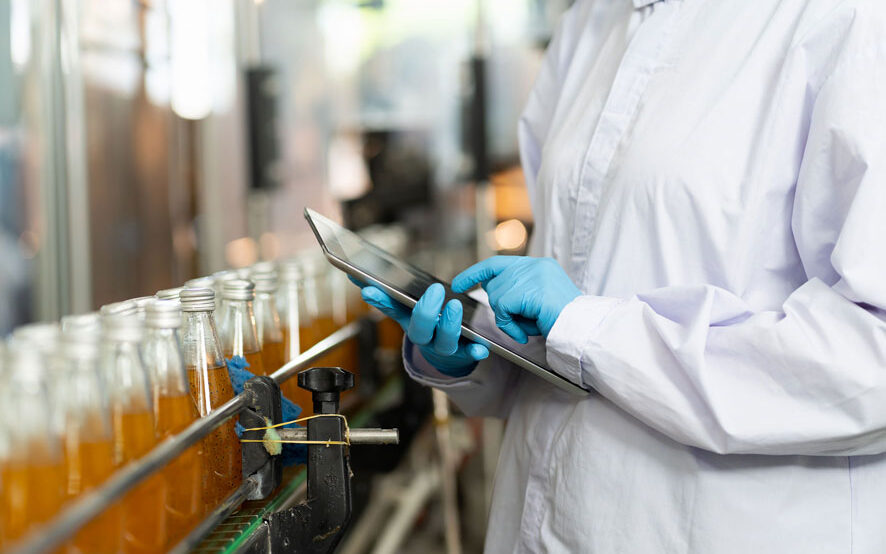

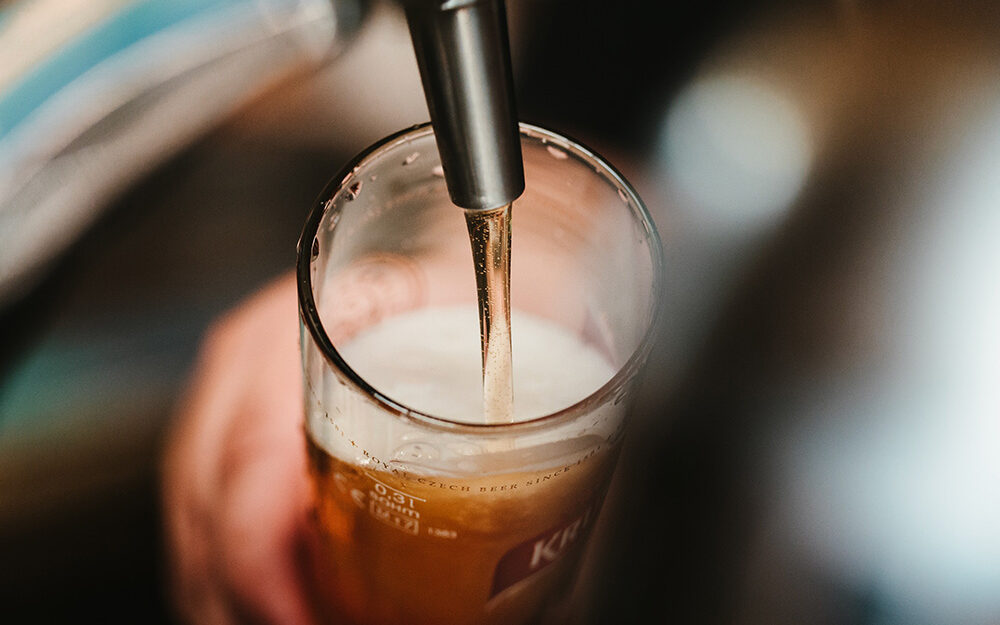
Food & Beverage
We help you maintain your quality control standards and give you the confidence of knowing exactly what is in your processes.

Probiotics
For QC control, our lab enumeration services help you confirm consistency of your probiotics from lot to lot. Monitor microbial communities, or try our phenotyping services to optimize your final product.
Microbial Identification
Biolog for Identification Services
For decades our lab has set the industry standards—we developed and refined many identification techniques that have become today’s standards.
ENUMERATION: TOTAL PLATE COUNT
Regarding food safety & probiotics, enumeration is essential for product quality.
Ensuring potency and accuracy can give you an edge over the competition. By offering enumeration testing or total plate count services (TPC), we ensure the bases are covered to determine viability and identity.

USP <797> Testing
Ensuring Compliance with Industry Standards
We assist you in achieving and maintaining compliance
with USP <797> standards, ensuring the safety and quality
of pharmaceutical and healthcare products.
– Bacterial enumeration and identification
– Fungal enumeration and identification
– Gloved fingertip bacterial sampling
– Media fill test
EXCITING NEW SERVICES
Biolog for Phenotyping Services
Not only has Biolog Lab Services moved to a shiny new lab in Newark, DE, but we are now offering phenotypic identification and analysis services.
We use our newest innovation, the Odin™ platform, combined with Phenotype MicroArrays will help you classify and differentiate metabolism and respiration from biomass production (if that’s important to you). We customize the experimental conditions for your specific strains and meticulously gather the data, which is summarized in a detailed report.
Our phenotyping services offer a classic and reliable method to characterize microbial strains. What’s more, with our multiomic approach, you can be confident in obtaining the precise identification you seek.

Resources
Advantages of multiomic microbial identification.
We cover multiomic identification: unveiling the identities of bacteria, yeast, and fungi from all angles.
Genotypic Analysis: Deciphering genetic codes with Sanger DNA Sequencing. Sequencing of ribosomal RNA regions of bacteria and fungi.
Proteotypic Profiling: Exploring protein landscapes through Bruker MALDI-TOF. Analysis of ribosomal proteins.
Phenotypic Insights: Odin’s lens into microbial behaviors. Analysis of biochemical reactions, acids and salt tolerance, metabolism, fermentation, etc.
We review Biolog’s solutions to identify thousands of aerobes, anaerobes, yeast, and filamentous fungi.
Biolog for Fungal Identification
Lab Services now offers a customized approach to challenging filamentous fungal ID.
With our novel method for subculturing, sample lysis and proprietary media that reduces contamination risk, we can now:
– effectively prepare the fungal samples
– ensure quick, effective, and accurate sample identification
– all in an average of 3 business days, or as fast as next day
Rapid Microbial Identification with MALDI-TOF Technology: Navigating the Fast Lane
In a world where speed, accuracy, and affordability rarely align, the emergence of MALDI-TOF technology has rewritten the rules for microbial identification. Join us for an insightful webinar as we delve into microbial identification, now revolutionized by MALDI-TOF. At Biolog Lab Services, we’ve seamlessly integrated this cutting-edge technology into our microbial identification and characterization suite. In this session, we’ll introduce you to the power of MALDI-TOF, highlighting its suitability for microbial identification, and shed light on how our implementation of this technique synergizes with other contamination identification and microbial classification methods.
Don’t miss this opportunity to explore the fast lane of microbial identification!




